Applications that make the cut
1 Feb 2019 by Evoluted New Media
There’s been a lot of news and hype about CRISPR, but what can it actually do? Dr Bahri Karacay take us through the hopes and limitations of the technique...
Unless you’ve been in total isolation for the last few years, you will have heard quite a bit about CRISPR. So, why all the fuss and what are we hoping it can do?
CRISPR-Cas9 is a gene editing system that has the scientific community pretty excited. But gene editing itself isn’t new, and in fact, neither is CRISPR. CRISPR wasn’t actually invented, it was discovered, or perhaps more accurately, noticed.
In 1987, a Japanese team of researchers studying E. coli were the first to notice CRISPR-associated repeat sequences in the bacterial genome but could not ascertain their significance. Others had identified similar clusters in other microbial species, and these sequences were ultimately termed Clustered Regularly Interspaced Short Palindromic Repeats or CRISPR. However, it wasn’t until 2007 that scientists were able to demonstrate that these clusters functioned as part of a bacterial immune system that protects against invading foreign DNA. CRISPR systems generally function by inserting short genetic sequences from foreign invading elements (viruses, etc.) into the genome, which are then used as a sort of genetic memory to detect and prevent from repeat infection. Just five years later this system was repurposed and exploited for precise genome editing in eukaryotic cells.
So how does CRISPR work as a gene editing system? When provided with a 20-nt guide RNA (gRNA) that matches a genomic target site, the Cas9-gRNA complex can then bind to this site and cut the DNA. The lesion is rapidly healed with native DNA repair machinery, which often results in a disruption of gene function (knock out). CRISPR can also be used to insert new DNA sequences, which introduced the possibility of therapeutic gene editing in the clinic.
The CRISPR-Cas9 system can potentially be used to cut any genome at virtually any place and it is both incredibly easy and inexpensive. It is therefore no surprise that CRISPR has become one of the most exciting new tools in biomedical research. To illustrate this, in 2011 there were less than 100 published papers on CRISPR, but this rose to more than 14,000 in the last year.
Specific and efficient CRISPR holds tremendous promise in many applications but key to all of this is its specificity and efficiency. Thus, major efforts have been devoted to improving these characteristics to a point where CRISPR-Cas9 genome editing can be performed accurately and safely, particularly as we approach the clinic.
There are many methods available for delivering CRISPR-Cas9 reagents into living cells. However, it is widely accepted that first binding Cas9 to a gRNA outside of cells (aka ribonucleoprotein or RNP complex), and then delivering the resulting complex directly into living cells provides the least opportunity for unwanted collateral damage.
Other delivery methods that rely on plasmid or viral vectors that maintain constant and long-lasting expression of CRISPR-Cas9 components. These methods are often prone to stimulation of the innate immune system and typically carry the biggest risk of off-target editing. A major benefit of the RNP delivery method is that CRISPR-Cas9 reagents are rapidly eliminated from cells which minimizes the opportunity for immune system stimulation and off-target editing. In other words, the RNP delivery method is akin to a stealth attack – it gets in, does the job accurately, and gets out quickly with as little detection as possible.
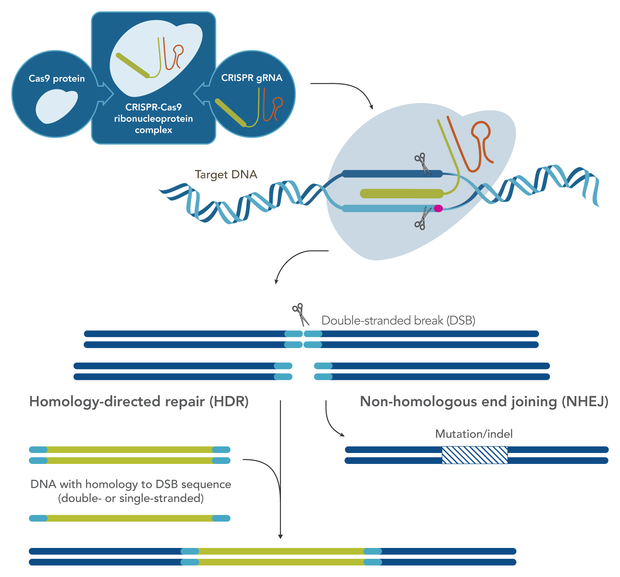
[caption id="attachment_70282" align="alignleft" width="372"] Figure 1. Effective CRISPR-Cas9 gene editing. Cas9 enzyme is delivered with gRNA as an RNP complex. gRNA directs Cas9 to excise the target DNA with a double stranded break. For a knock-out, the cell repairs the break by NHEJ. For a knock-in, the desired insert is delivered as a DNA donor template that has homologous sequences with either end of the break to stimulate HDR.[/caption]
Figure 1. Effective CRISPR-Cas9 gene editing. Cas9 enzyme is delivered with gRNA as an RNP complex. gRNA directs Cas9 to excise the target DNA with a double stranded break. For a knock-out, the cell repairs the break by NHEJ. For a knock-in, the desired insert is delivered as a DNA donor template that has homologous sequences with either end of the break to stimulate HDR.[/caption]
Once a cut has been made, two of the most common repair outcomes include the direct sealing of blunt ends via the non-homologous end-joining pathway (NHEJ), or homologous recombination by a homology-directed repair (HDR) pathway in the presence of a suitable donor template (Figure 1).
The NHEJ pathway typically seals CRISPR-Cas9 cuts by an error-prone process that results in an insertion or deletion (Indel) of one to several DNA bases. HDR, on the other hand, requires the addition of a donor template DNA (the sequence to be inserted) that contains “homology arms” that overlap with each end of the break and typically provides a more “perfect” or error-free repair. As HDR is more reliable, it is generally speaking the preferred option for knocking in sequences and achieving reliable insertions while NHEJ is more often used for knock-out experiments.
So how is CRISPR being used and what are the hopes for the future?
Research models CRISPR-Cas9 is an obvious choice to advance the creation of traditional (global), or conditional knockout and transgenic animal models, in order to speed up our understanding of what different genes in different organisms do.
Cre-LoxP conditional knockout mouse models where a gene is flanked by LoxP sites (floxed), are among the most useful genetically engineered models in biomedical research. Addition of the Lox sequences enables genes to be activated, repressed, or even exchanged at a developmental stage or in specific tissues. Therefore, floxed mouse models provide the opportunity to define essential gene functions in both global and tissue-specific contexts, and are particularly critical for analysing genes that have essential functions in early development.
CRISPR has been extensively used to generate mice models with small insertions and deletions, however it has been difficult to obtain efficient and accurate insertion of long sequences. Recently, a new method called Easi-CRISPR (Efficient Additions with SsDNA Inserts-CRISPR) has been developed that overcomes this difficulty, enabling and simplifying the creation of mice carrying conditional and insertional alleles for large-scale floxed animal resources.1 The method enables insertion of long single-stranded DNA donors with a preassembled RNP complex into cells for efficient HDR-mediated knock-in, applicable not only in Cre-LoxP animal resources, but also across many low- and high-throughput genome engineering applications.
In addition to knock-out and knock-in approaches, CRISPR technology has also been explored as a tool for activating or silencing gene expression to further our understanding of genes and their biological pathways.
Agricultural biotech CRISPR-Cas9 also presents an efficient way to develop desirable traits in agriculturally important crops – particularly where traditional techniques are difficult. Potato is one such example. Cultivated potato is one of a number of crops that are tetraploid, which complicates research and breeding via crossbreeding. Therefore, researchers have been using CRISPR-Cas9 to study and develop commercially important traits, but the method still requires optimising for efficiency and specificity to be able to safely add specific traits.
CRISPR methods, in theory, have the potential to correct a myriad of disorders – although we may be some way off seeing it become a routine therapeutic method on the NHSRecent studies demonstrate that delivery of CRISPR reagents in RNP format is a promising alternative to plasmid-based delivery method for potato breeding. RNP delivery provides high editing efficiency in developing new potato lines without unintended DNA integration.2
This year, however, the EU Court of Justice ruled that CRISPR-Cas9 gene-edited plants should be subject to the same regulations as genetically modified organisms. While there is still significant potential for use in plant research, the ruling has dampened enthusiasm for the application of the technology to develop commercial traits. However, if CRISPR is proven as safe as the methods that are already exempt from the law, it could be granted an exemption. The work being done to increase specificity and efficiency, such as RNP complex delivery, may help us to reach that milestone.
Human therapeutics One of the most exciting opportunities for CRISPR-Cas9 genome editing is its use as a therapeutic tool for the treatment of human diseases. CRISPR methods, in theory, have the potential to correct a myriad of genetic disorders. Although we may be some way off from CRISPR becoming a routine therapeutic method on the NHS, there is a lot of encouraging work in the right direction. Some of the biggest barriers to CRISPR-Cas9 in human therapeutics is successful delivery to the intended tissues and its safety – here defined as specificity and efficiency. We must ensure that editing is highly specific with minimal off-target editing events, and if specific off-target editing events are unavoidable, it must be demonstrated that they do not cause disease.
Despite these issues, there have recently been some great leaps forward in ex-vivo genome editing – taking the cells out of the body, editing, and then replacing them to correct the disease phenotype. This circumvents the problem of reliably targeting nucleic acid and protein-based therapies to a wide variety of tissues and cell type in the human body – something which at present, we are unable to do.
In a recent paper by Vakulskas and Dever et al. in Nature Medicine, researchers at Integrated DNA Technologies (IDT) in collaboration with Stanford University introduced a novel high-fidelity Cas9 enzyme (HiFi Cas9) that when delivered with the RNP format has greatly improved specificity and excellent on-target potency. This is particularly important as previous attempts to increase specificity had frequently sacrificed good overall editing efficiency. It was demonstrated that the HiFi Cas9 enzyme allowed for correction of the sickle cell disease-causing mutation in human hematopoietic stem and progenitor cells (HSPCs) – with off-target editing below the limits of detection.3 The work is even more significant as HSPCs are notoriously difficult to edit.
Several additional strategies are being pursued to use CRISPR in the development of modified chimeric antigen receptor (CAR) T-cell therapies for the treatment of various cancers. T lymphocytes are essential regulators and effectors of adaptive immune responses and the study of gene function in primary T-cells is highly relevant for T-cell based immunotherapies. Previous attempts to use CRISPR-Cas9 gene editing in primary T-cells were very variable and also needed T-cell receptor (TCR) stimulation, which meant that genes involved in activation or differentiation could not be studied. Recent work however, has described an optimised approach for effective Cas9 RNP transfection in both primary human and mouse T-cells without the need for TCR stimulation. This method will likely greatly enhance target gene discovery and validation in primary T-cells and simplify the editing process for next-generation immunotherapies.4
This work shows that current RNP methods together with HiFi Cas9 enzymes hold great promise for achieving the level of efficiency and specificity for successful ex vivo therapeutics applicable in many human diseases.
CRISPR of the future As well as the aforementioned CRISPR applications there are also some remarkably innovative and unexpected uses for this system. For example, CRISPR has been combined with NGS to create an ultra-accurate sequencing strategy for very high on-target enrichment. The method termed CRISPR-DS employs CRISPR to digest fragments of a predetermined length, enabling size selection before library preparation.5 Another innovative approach employs CRISPR as a method to tag endogenous proteins with a luminescent peptide. By using tagged-CRISPR instead of inserting recombinant reporter genes and overexpressing them, biological processes can be studied in their native contexts.6
The big question now is whether CRISPR will live up to all the hype. We’ve made tremendous progress in the last few years, and with all the current work being done, it seems almost certain to do so. However, expectations should be tempered with timeframe considerations. There is often much excitement when new technologies emerge, and an expectation that breakthroughs e.g. new medical therapies will quickly follow. Doubt sets in when this doesn’t happen as quickly as anticipated – therapeutic monoclonal antibodies are a good example. It took more than a decade to begin developing these as medical therapies, but today monoclonal antibodies are some of the most exciting new therapeutics. This is also true of antisense, now employed in a number of successful drugs in late-stage clinical trials, and RNA interference—discovered in 1998— employed for the first time in an approved medical therapy just this year.
CRISPR is already having a massive impact and will continue to do so in several fields including biomedical research, agricultural biotechnology, and medicine. However, we might need to wait a while longer to see it safely introduced into medical therapeutics. It is increasingly likely though that ex vivo RNP-mediated therapeutic editing of HSPCs and T-cells will be some of the first CRISPR-Cas9 programmes to enter clinical trials within the next few years. And while safety and efficacy can only really be tested in phase I human clinical trials, it will be vital to use the best possible reagents – such as HiFi Cas9 enzymes and chemically-modified gRNAs – for the best possible specificity and efficiency to maximise safety and minimise risks.
References
- Quadros RM, et al. (2017) Easi-CRISPR: a robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biology, 18(1):92.
- Andersson M, et al. (2018) Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol Plant.
- Vakulskas, CA, et al. (2018). A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nature Medicine 24(8), 1216-1224.
- Seki A, Rutz S. (2018) Optimized RNP transfection for highly efficient CRISPR/Cas9-mediated gene knockout in primary T cells. J Exp Med 215(3):985-997.
- Nachmanson D, et al. (2017) CRISPR-DS: An efficient, low DNA input method for ultra-accurate sequencing. bioRxiv. doi: 10.1101/207027.
- Schwinn MK, et al. (2017) CRISPR-mediated tagging of endogenous proteins with a luminescent peptide. ACS Chem Biol, 13(2):467-474.
 Dr Bahri Karacay is a Scientific Writer at Integrated DNA Technologies.
Dr Bahri Karacay is a Scientific Writer at Integrated DNA Technologies.