Understanding Nature’s scalpel
15 Dec 2016 by Evoluted New Media
The CRISPR-Cas system is already one of our most important genetic techniques – and it is one honed by evolution not humanity. So why did it evolve, how exactly does it work and what can we do with it?
The CRISPR-Cas system is already one of our most important genetic techniques – and it is one honed by evolution not humanity. So why did it evolve, how exactly does it work and what can we do with it? Hannah Koslowski gives us a backgrounder on the sharpest of genetic tools…
Recently, the field of genetic engineering has exploded. The CRISPR/Cas system has become the leading mechanism in genome engineering in just a few years since its discovery. This system has been adapted for many purposes, such as determining the functions of genes, curing diseases, modifying crops, and tracking outbreaks.
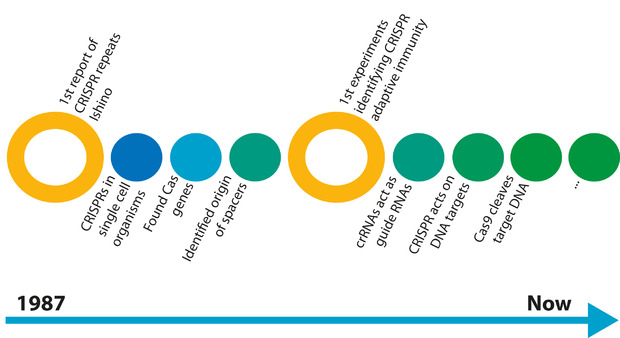
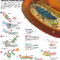
The first mention of the CRISPR/Cas system was in 1987 when Dr Yoshizuma Ishino mentioned that he found strange DNA repeat sequences in the bacteria he was studying¹ (figure 1). There was no follow up to this discovery until the year 2005, when three different groups reported that the characteristic “spacer” sequences were almost identical to viral genomic sequences²-?. Shortly after this discovery, another group reported that this CRISPR/Cas system may function as a primitive immune system in bacteria?. By this point, the scientific community was buzzing and the race to understand the CRISPR/Cas system was on. When a DNA virus infects a cell, the virus injects its DNA genome into the cell and, in some cases, the viral genome is integrated into the host genome. In multicellular organisms, the infected cell is detected then killed during an immune response. However, being single-celled organisms, bacteria don’t have an immune system and must deal with the virus itself, or face death.
[caption id="attachment_56806" align="alignnone" width="620"]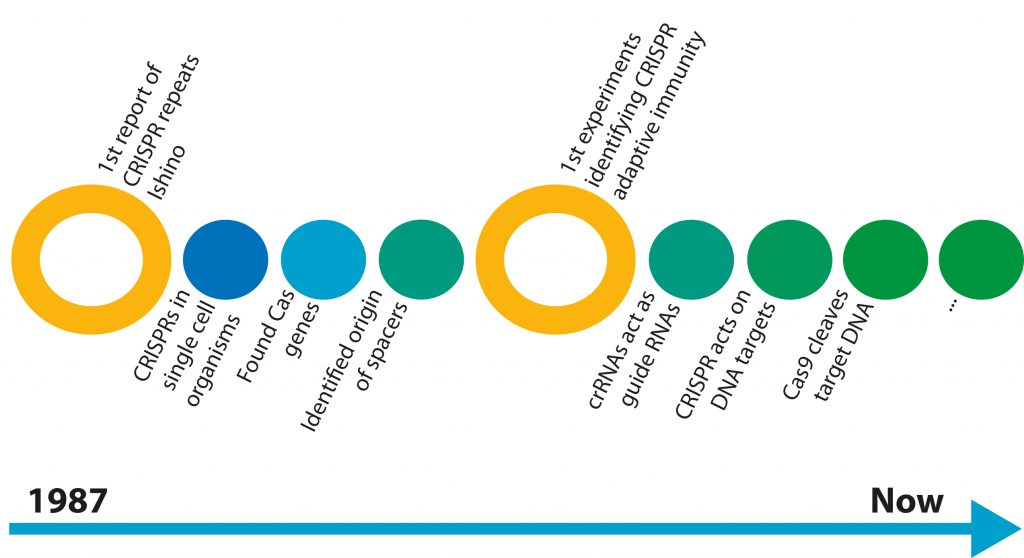 Figure 1: The elucidation of CRISPR and Cas9.[/caption]
Figure 1: The elucidation of CRISPR and Cas9.[/caption]
And while viruses are evolving to overcome bacterial defences – bacteria have many mechanisms in place to combat viruses and prevent infection. This billion year old war between bacteria and viruses is often called the “bacteria/phage arms race,” whereby each party tries to get the upper hand in order to survive?,?. But bacteria, as simple as they are, have developed a spectacular way to combat these bacteriophage (a virus that infects bacterial cells) invasions: the CRISPR/Cas system. This system allows the bacterium to identify DNA invaders and remove the threat before any damage can be done.
Simply put, the CRISPR/Cas system is a bacterial immune system, which allows a bacterium’s progeny to remember foreign invaders, thus preventing future infections. To understand the mechanism of the CRISPR/Cas system, it is vital to understand some basic genetics. Every organism has a genetic code made up of DNA that is "transcribed" to RNA (there are numerous different types), which is then recoded ("translated") into protein. Recall that the genetic code encodes all proteins and their products in an organism; it is used to explain gene expression, protein synthesis and organismal functioning as a whole. The CRISPR/Cas system can edit and rewrite the genetic code, the foundation of all organisms.
Simply put, the CRISPR/Cas system is a bacterial immune system, which allows a bacterium’s progeny to remember foreign invaders, thus preventing future infections.
There are three different CRISPR/Cas systems but here we will only delve into the most common system used in scientific research: the CRISPR II system. The term CRISPR stands for Clustered Regularly Interspaced Short Palindromic Repeats, which indicates that it is a portion of DNA that has multiple repeat sequences (direct repeats), separated by variable sequences (spacers). The spacers are DNA segments that match the DNA of bacteriophages; this way, CRISPR enables the bacterium to create its own pathogen library.
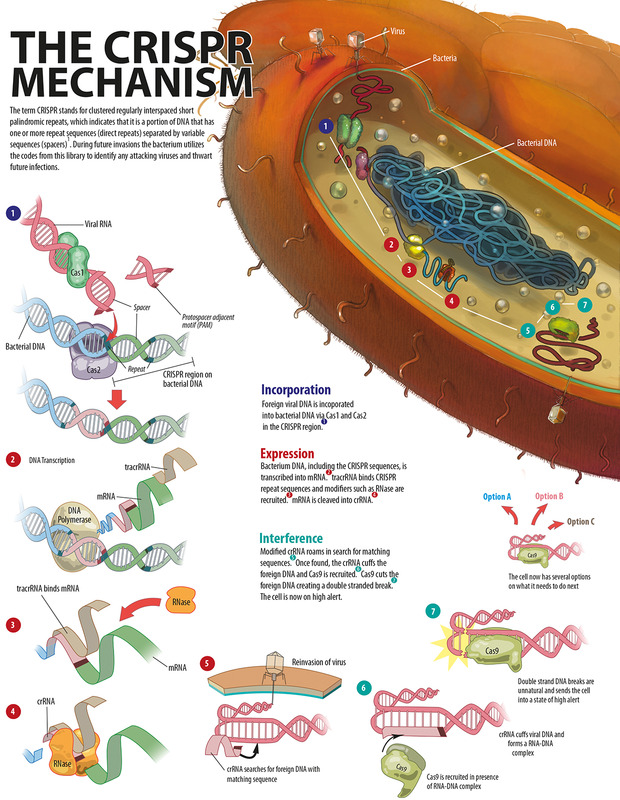
This library contains DNA sequences from different strains and species of phages that have previously infected that bacterium or that have infected the ancestors of that bacterium; all archived in the bacterial cell’s genome. The process of cataloguing all the “books” in the CRISPR library is referred to as the acquisition stage (figure 2). During future invasions, the bacterium utilises the codes from this library to identify attacking viruses and thwart future infections. Like all successful systems, the CRISPR/Cas system needs to be regulated. These regulators prevent the system from losing control. In the CRISPR/Cas system, protospacer adjacent motifs (PAMs) prevent random cleaving of the bacterial genome, therefore protecting the bacterium from destruction by CRISPR/Cas?,?. The PAMs are DNA sequences – only found in the invading DNA and not the bacterial genome – that mark the ends of target sequences. Furthermore PAMs do two things by marking the cleavage sites: facilitate sequence specific cleavage of the invader and prevent random cleavage of the bacterial genome. Without proper PAM recognition by Cas9, a DNA cleaving protein, the foreign viral DNA cannot be labelled as an invader.[caption id="attachment_56804" align="alignnone" width="620"]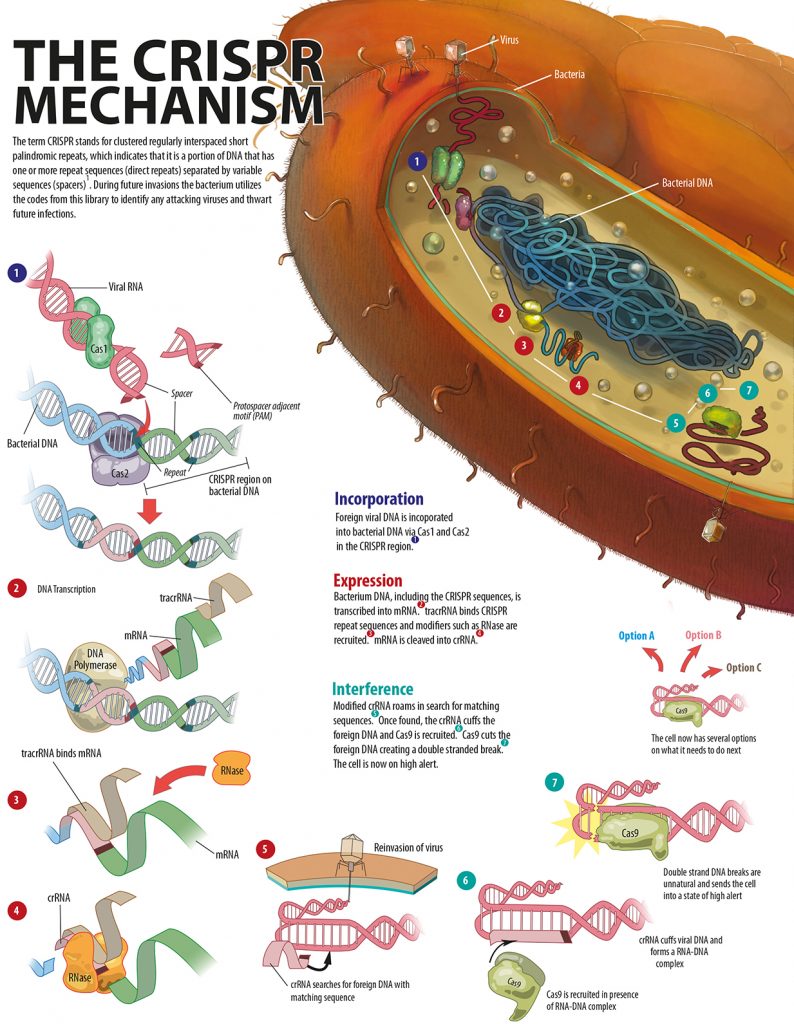 Figure 2. An explanation of the CRISPR/Cas mechanism.[/caption]
Figure 2. An explanation of the CRISPR/Cas mechanism.[/caption]
Using the information from the CRISPR/pathogen library – all found within the bacterial genome –each of the spacers are cleaved into small fragments. The small fragments are transcribed into CRISPR RNAs (crRNAs) and transactivating CRISPR RNAs (tracrRNAs)¹?. The tracrRNAs aid in the processing of crRNAs, which are short sequences specially designed to bind and match the archived phage DNA sequences. However, in scientific experiments the crRNA and tracrRNA are fused into one molecule called the single guide RNA (sgRNA), which has the functional capabilities of both components¹¹. The cleaved crRNAs are released within the cell where their function is to search for matching sequences and then recruit Cas9. The crRNAs act as detectives: once they catch their culprit, they "cuff him". After a match is made between the crRNA and the foreign DNA, the two strands bind together. The "cuffed" DNA sequence has now been flagged as an invader.
The invader DNA is now the target for Cas9. It is also the job of the crRNA to lead Cas9 to the invading pathogen DNA¹². Once the Cas9-crRNA complex reaches the “scene of the crime”, Cas9 cleaves both strands of DNA precisely at the edge of the PAM region in the crRNA-pathogenic DNA complex. Cas9 cleaves both DNA strands causing a double stranded break in the invading DNA – both DNA strands are cut¹². The double stranded DNA break is pivotal in the CRISPR/Cas system and acts to inactivate the viral genome. However, if a double stranded break occurs in the cell’s own genome, the cell has several options, the most common being:
- The break is fixed by homologous recombination. In this process, also cited as homology directed repair, other enzymes are recruited to the site of the double stranded break to patch the break using a DNA template. This template is a sequence of DNA – which can be designed and supplied to the cells by scientists – that will be used as a guide for repair processes¹³,¹?.
- The break is fixed by non-homologous end joining. In this process there is no DNA template available. While sometimes the two DNA ends are glued back together, other times the final DNA sequence is altered, which may lead to gene inactivation¹?. These pathways show that by using the CRISPR/Cas system, either the bacterium or scientist can induce many different changes in the cell. Obviously, some pathways are more favourable for certain applications, but the ability to edit DNA in a targeted simplistic manner is a much needed gift for the future of scientific research.
The CRISPR/Cas system may have started as a bacterial defence mechanism against foreign pathogens, but humans have repurposed this unusual defence system because it is a system that can be used to edit any DNA sequence. There are uses for CRISPR in genetic engineering, drug therapies, outbreak analysis and much more. In this section, we will explore the current uses and future directions of the CRISPR/Cas system. Scientists use CRISPR to rapidly modify the genome and create models of disease. CRISPR can easily be used to find the functions of specific genes by turning the target gene “off” and then identifying what has changed in the cell or organism. Similarly, CRISPR is used to mimic genetic mutations linked to specific diseases therefore allowing the scientist to study the effects of these modifications on disease development, and the application of potential disease treatments. This is done by introducing one or several mutations – implicated in a naturally occurring human disease – in the DNA of a model organism, such as a mouse, using CRISPR/Cas.
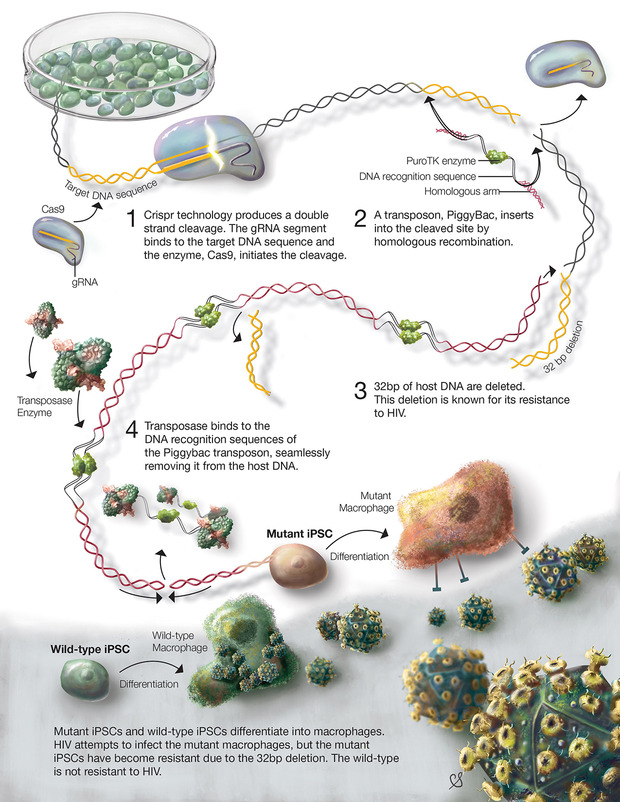
About 80% of genetic diseases are caused by mutations, spontaneous or inherited, that change only a small segment of the genetic code which can influence disease expression or severity¹?. Currently, there are many drugs that may decrease symptoms or prolong onset of some diseases, but cures are far and few between. The surest way to cure a genetic disease is to correct the disease causing mutation. For example, CRISPR/Cas has been used to induce resistance to HIV infection (figure 3)¹?. HIV is a deadly virus, which infects human immune cells, such as T cells and macrophages. Most individuals with HIV, experience disease progression and develop AIDS which ultimately leads to death. Unfortunately, there is no cure for HIV, but recent research using CRISPR/Cas aimed to prevent HIV infection. In these experiments, the CD4 receptor – used by HIV to infect cells – was modified in human stem cells using the CRISPR/Cas system¹?. The stem cells were transformed into macrophages and were able to resist HIV infection.[caption id="attachment_56805" align="alignnone" width="620"]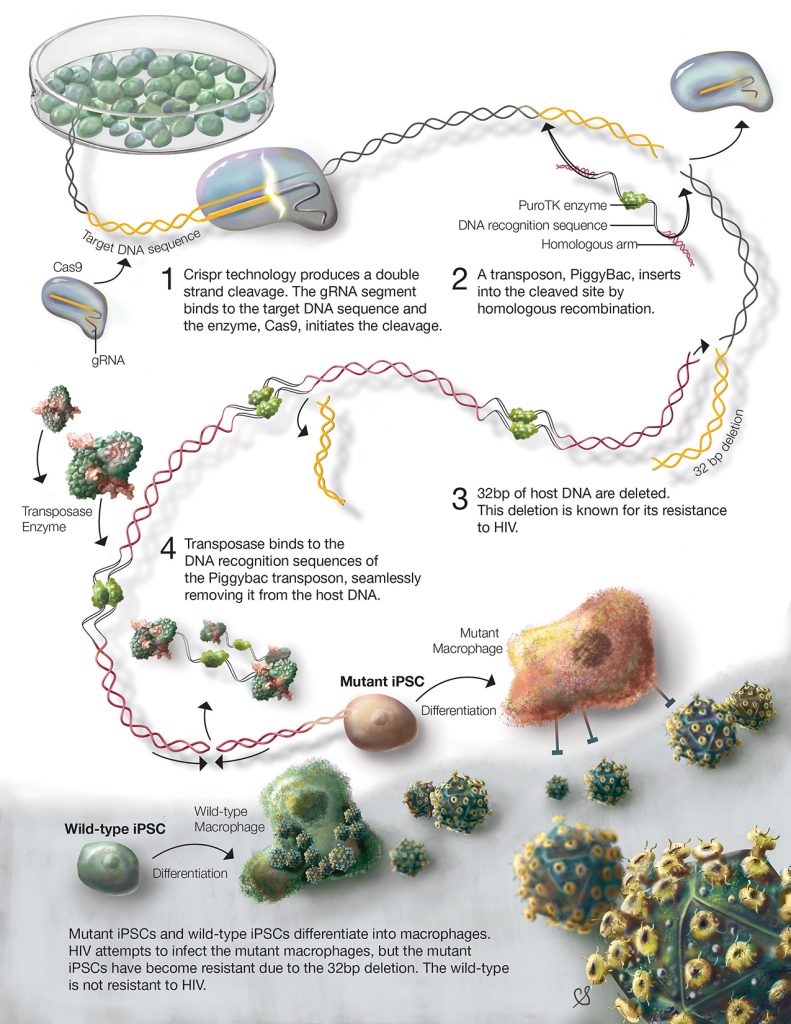 Figure 3. The process of modifying stem cells using CRISPR/Cas to induce HIV-1 resistance. bp indicated base pairs (fundamental DNA units)[/caption]
Figure 3. The process of modifying stem cells using CRISPR/Cas to induce HIV-1 resistance. bp indicated base pairs (fundamental DNA units)[/caption]
Assuming we can use CRISPR/Cas to cure patients, we need to contemplate the ethical implications of these treatments. Even though the CRISPR/Cas system can be used to eliminate some diseases, it has recently gained copious media attention and ethics-related criticism for its ability to change the human genome. Some of the concerns are associated with potential non-therapeutic uses, which originated when a group of scientists edited the genome of “non-viable” human embryos¹18,¹19. Scientists are also concerned that those types of studies will foster prejudice and impact their ability to conduct seminal studies looking to cure or understand genetic diseases. Currently, there is discussion about regulating the cell types and animals that could be modified using CRISPR/Cas, therefore increasing patient safety and preventing public outcry¹?.
The CRISPR/Cas system may have started as a bacterial defence mechanism against foreign pathogens, but humans have repurposed this unusual defence system because it is a system that can be used to edit any DNA sequence.
CRISPR/Cas has also been used for phage resistance in dairy culture. The dairy industry is a multi-billion dollar industry that uses bacteria to complete the fermentation process, which leads to the thickening of milk. Phages often contaminate raw milk since they can survive some pasteurisation and sanitation procedures and thereby infect and kill important dairy microbes, costing companies millions of dollars. To combat this problem, sustainable bacterial cultures were developed using CRISPR/Cas to ensure that bacteria can resist attack from phages. In this case, bacteria are infected with a bacteriophage that commonly affects dairy cultures. The bacteria that survive this attack will harbour a CRISPR spacer that targets that phage DNA therefore making it resistant to infection from that virus and suitable for dairy production²?. This bacterial culture prevents both revenue loss and food shortage.
Furthermore, some of the most interesting uses for CRISPR/Cas do not rely on its cleaving ability. Instead, new uses for the CRISPR/Cas system have a deactivated Cas9 (dCas9) domain so that it cannot cut DNA or the system requires only the sgRNAs. This is due to precise ability of the CRISPR/Cas system to accurately target a DNA sequence of interest; deactivating Cas9 does not affect the crRNA's ability to identify the matching DNA sequences. Using dCas9 scientists have been able to search the entire genome for various purposes. In select experiments, scientists attach a fluorescent marker to the dCas9; therefore the dCas9 complex can be tracked within the cell with the use of microscopes²¹.This allows scientists to see the location of the target sequence in the cell and its movement over time – live cell imaging. The location of a gene can tell scientists about the activity of that gene in a cell and if the location of the gene changes²²,²³. Furthermore, scientists now think that the location of a gene in a cancer cell may give us information about the aggressive nature of tumors, susceptibility to treatment, and improved diagnosis²².
Similarly, the identification of bacterial strains using CRISPR spacer sequences can be used as a tool to track the evolution of bacteria.
Epigenetics involves modifications to the DNA that do not affect the DNA sequence. These modifications affect the conformation or structure of the DNA, gene expression or affect the types of proteins that associate with that DNA sequence²?. Additionally, it was found that certain DNA modifications can turn a gene “on”, therefore activating it. Scientists can add an enzyme – which modifies the molecules attached to the DNA – to dCas9 to see whether this changes expression of that gene or cell function²?. Most importantly, epigenetic mutations have been associated with various cancers, leukemia, mental retardation and several other diseases and by using CRISPR/Cas, scientists can move the field of epigenetics forward, and gain a better understanding of the mechanisms of disease²?.
Another interesting use for the CRISPR/Cas system is in outbreak analysis. This is employed when there are an unusually large number of individuals infected with a given microorganism at the same time or in the same area. In an outbreak, health care workers need to pinpoint the origin of the outbreak, and the bacterial strain involved. A CRISPR/Cas identification system can be used to differentiate between bacterial subtypes within a serotype. A serotype is a subgroup within a species of microorganism that have the same cell surface structures²?, while a subtype is a genetic variant of a microorganism.
Similarly, the identification of bacterial strains using CRISPR spacer sequences can be used as a tool to track the evolution of bacteria. As we already know, the CRISPR region of the genome changes due to the acquisition and loss of spacers. This process shows the evolution of the bacterium. A novel use of the CRISPR system is in the study of fruit bacteria. Certain bacterial pathogens can cause diseases in apples, pears and many other fruits and CRISPR/Cas can be used to identify difference between bacteria infecting different fruits or the same fruits in different regions²?.
The CRISPR/Cas system is radically changing the world around us. It appears that the CRISPR/Cas system could be manipulated to solve many cell-related problems; not just in humans, but in fruits and dairy cultures and in other living organisms. The two most valuable characteristics of the CRISPR/Cas system are the ability to bind specific gene sequences and edit the target gene; both actions are completed with extraordinary efficiency. As we develop new uses for CRISPR/Cas there is no telling what efficiencies could be made and information discovered. Science is always looking to push the boundaries, and now CRISPR/Cas allows us to push harder and faster.
Author: Hannah Koslowski is a PhD student at the University of Toronto. Hannah works closely with Dr Alberto Martin, an established immunologist at the University of Toronto.
Acknowledgements: We would like to thank Erina He, Sonia Seto and Caitlin Swanberg for the time they took to develop the graphics used in this article.
References: 1. Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987 Dec;169(12):5429-33. 2. Bolotin A, Quinquis B, Sorokin A, Ehrlich SD. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology. 2005 Aug;151(Pt 8):2551-61. 3. Mojica FJ, Díez-Villaseñor C, García-Martínez J, Soria E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol. 2005 Feb;60(2):174-82. 4. Pourcel C, Salvignol G, Vergnaud G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additionaltools for evolutionary studies. Microbiology. 2005 Mar;151(Pt 3):653-63. 5. Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV. A putative RNAinterference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct. 2006 Mar 16;1:7. 6. Stern A, Sorek R. The phage-host arms race: shaping the evolution of microbes. Bioessays. 2011 Jan;33(1):43-51. Review. 7. Marraffini LA. CRISPR-Cas immunity against phages: its effects on the evolution and survival of bacterial pathogens. PLoS Pathog. 2013;9(12):e1003765. 8. Jore MM, Brouns SJ, van der Oost J. RNA in defense: CRISPRs protect prokaryotes against mobile genetic elements. Cold Spring Harb Perspect Biol. 2012 Jun 1;4(6). pii: a003657. 9. Sashital DG, Wiedenheft B, Doudna JA. Mechanism of foreign DNA selection in a bacterial adaptive immune system. Mol Cell. 2012 Jun 8;46(5):606-15. Kozlowsk iHN Page 12. 10. Pougach KS, Lopatina AV, Severinov KV.CRISPR Adaptive Immunity Systems of Prokaryotes. Mol Biol. 2012 Apr; 46 (2): 175-182. 11. Terns RM, Terns MP. CRISPR-based technologies: prokaryotic defense weapons repurposed. Trends Genet. 2014 Mar;30(3):111-8. 12. Liu L, Fan XD. CRISPR-Cas system: a powerful tool for genome engineering. Plant Mol Biol. 2014 Jun;85(3):209-18. 13. Helleday T, Lo J, van Gent DC, Engelward BP. DNA double-strand break repair: from mechanistic understanding to cancer treatment. DNA Repair (Amst). 2007 Jul 1;6(7):92335. Review. 14. Lees-Miller SP, Meek K. Repair of DNA double strand breaks by non-homologous end joining. Biochimie. 2003 Nov;85(11):1161-73. Review. 15. Pelletier S, Gingras S, Green DR. Mouse genome engineering via CRISPR-Cas9 for study of immune function. Immunity. 2015 Jan 20;42(1):18-27. 16. Niu J, Zhang B, Chen H. Applications of TALENs and CRISPR/Cas9 in human cells and their potentials for gene therapy. Mol Biotechnol. 2014 Aug;56(8):681-8. 17. Ye L, Wang J, Beyer AI, Teque F, Cradick TJ, Qi Z, Chang JC, Bao G, Muench MO, Yu J, Levy JA, Kan YW. Seamless modification of wild-type induced pluripotent stem cells to the natural CCR5?32 mutation confers resistance to HIV infection. Proc Natl Acad Sci U S A. 2014 Jul 1;111(26):9591-6. 18. Evitt NH, Mascharak S, Altman RB. Human Germline CRISPR-Cas Modification: Toward a Regulatory Framework. Am J Bioeth. 2015 Dec;15(12):25-9. 19. Cyranoski D, Reardon S. Chinese scientists genetically modify human embryos. Nature News. 2015 Apr. 20. Barrangou R, Horvath P. CRISPR: new horizons in phage resistance and strain identification. Annu Rev Food Sci Technol. 2012;3:143-62. K o z l o w s k i H N P a g e | 13 21. Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014 Jun 5;157(6):1262-78 22. Meaburn KJ, Gudla PR, Khan S, Lockett SJ, Misteli T. Disease-specific gene repositioning in breast cancer. The Journal of Cell Biology. 2009;187(6):801-812. doi:10.1083/jcb.200909127 23. Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, Park J, Blackburn EH, Weissman JS, Qi LS, Huang B. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell. 2013 Dec 19;155(7):1479-91. 24. Egger G, Liang G, Aparicio A, Jones PA. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004 May 27;429(6990):457-63. Review. 25. CDC. Serotypes and the Importance of Serotyping Salmonella. 2015 Mar.